Mahavir Singh’s lab seeks to unravel mysteries behind complex protein-nucleic acid interactions

Mahavir Singh likes to experiment, both in the lab at work and in the kitchen at home. When he tries out new recipes for his children to taste, he sometimes feels like a contestant on MasterChef. “They are very hard to please,” he says. When his cooking experiments fail, Mahavir uses it as an opportunity to educate his children about how research in the lab is fraught with similar challenges. “I give them an analogy from work. If we do an experiment 10 times, and if it works one or two times, I will call that a success, because 10-20% success is quite good in the initial stage when you are trying out new things, like a new recipe in the kitchen.”
Growing up in the hilly state of Uttarakhand, Mahavir wasn’t always sure about pursuing a career in research. After finishing high school in Delhi, he was initially interested in medicine. “I didn’t get a good enough rank. I did get admission to dental college, but I was very sure that I wouldn’t want to be a dentist.” He decided to pursue a Bachelor’s in biochemistry at Deshbandhu College of Delhi University, where he first got a taste for research, and developed an interest in visualising molecules.
After completing a Master’s in biochemistry in 2001 from Delhi University, he decided to go abroad for a PhD. “But going to the USA became a little difficult because of 9/11,” he remembers. He turned his sights to Europe and went to the Max Planck Institute of Biochemistry in Martinsried, Germany, to work with biochemist Tad Holak. “I had a great time there. We were working on understanding the cell cycle,” he recalls. His research focused on teasing out the structure of a cell cycle regulator called the retinoblastoma protein, which allowed him to gain hands-on experience in methods such as X-ray crystallography and NMR spectroscopy.
After his PhD, he pursued a postdoc under Juli Feigon, Professor in the Department of Biochemistry at the University of California Los Angeles (UCLA), USA, whose lab was trying to understand how proteins interact with DNA and RNA. Mahavir focused his attention on an enzyme called telomerase, which maintains the ends of the chromosome – the bundle of DNA and proteins that carries our genetic blueprint. Understanding telomerase structure and how it works is important because of its role in controlling ageing as well as allowing cancer cells to proliferate, Mahavir explains. “People want to find a molecule that can inhibit telomerase activity in cancer cells.”
When Mahavir left UCLA to join IISc as an assistant professor, his interest in the chromosome continued. His lab set out to understand how the end stretches of the chromosome, called telomeres, are remodelled. Under normal conditions, these straight ends are twisted into loops or complex structures called quadruplexes to keep them safe. When the DNA needs to be copied (replicated), these loops and structures need to be unravelled, and looped back again once replication is over. “The story is more complex because you have a set of proteins also bound to these structures,” adds Mahavir.
His lab is trying to pinpoint the role of proteins involved in telomere DNA remodelling. One protein in particular that caught their attention was hnRNPA1. A few years ago, his lab found that this protein has a “disordered” region that hones in on specific telomere DNA and RNA quadruplex structures, and binds to them, allowing the protein to unravel these structures. “That was the work of my first graduate student, Meenakshi Ghosh,” he says. Understanding the role this protein plays is critical, he adds, because there are several diseases linked to its malfunction, such as Amyotrophic Lateral Sclerosis (ALS), a debilitating condition that progressively destroys patients’ motor control, and has no cure.
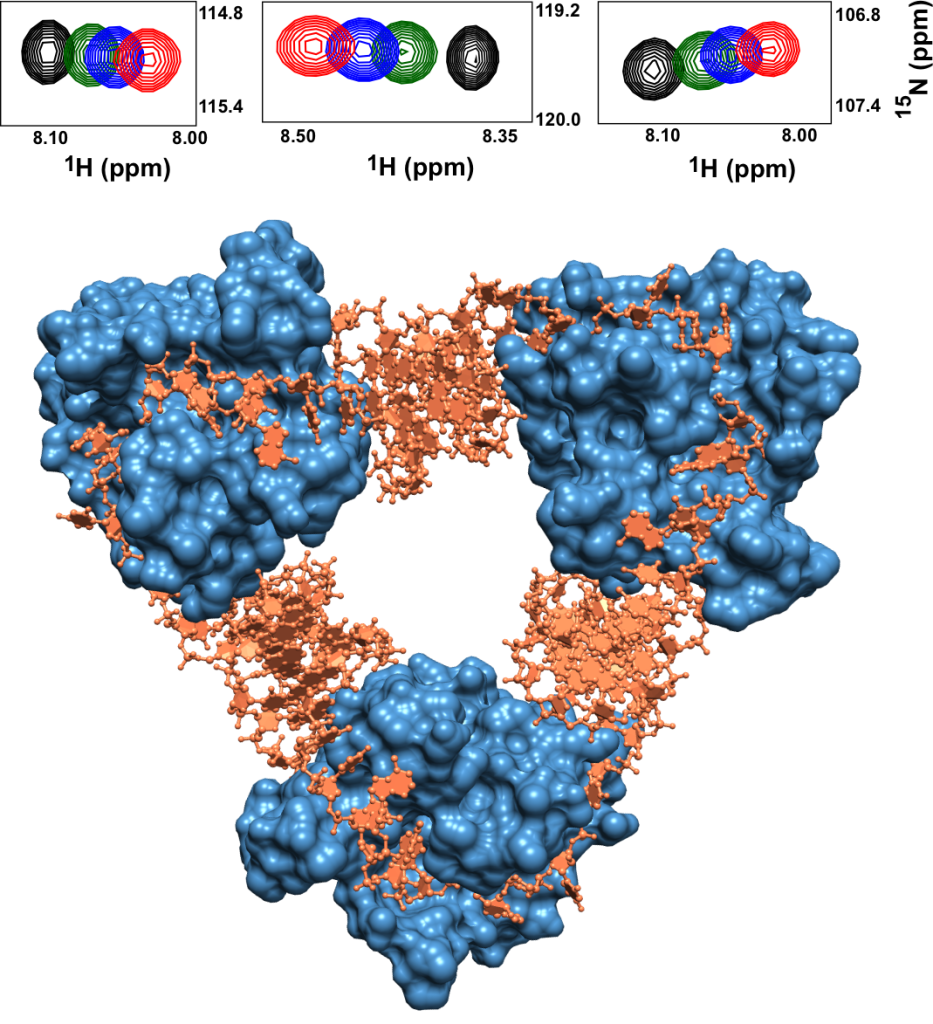
To study such protein-nucleic acid interactions, Mahavir’s lab uses a combination of biophysical and spectroscopic techniques. The most versatile among these is Nuclear Magnetic Resonance (NMR) spectroscopy, which leverages the magnetic properties of atoms to reconstruct protein structures. NMR spectra can give a readout for each amino acid residue in a protein, almost like a blueprint. When the protein binds to DNA or RNA, the corresponding shifts in the spectra can allow biophysicists like Mahavir to pinpoint which residues are doing the binding and to which form of DNA or RNA. “We can also quantify what is the strength of the binding,” he adds.
Soon after he joined IISc, Mahavir received funding under a Centre of Excellence grant from the Department of Biotechnology (DBT), Government of India, which allowed him to shift gears a bit and start investigating an interesting defence mechanism that certain bacteria adopt to fight viruses, called the toxin-antitoxin (TA) system. “Half of my lab is now working on this,” he explains.
When some viruses attack bacteria, they hijack the latter’s DNA, forcing the bacteria to spit out multiple copies of the virus, which burst the bacterial cell open, killing the bacterium and infecting others. To stop the virus from spreading, some bacteria kill themselves as an act of altruism. One of the key components in this process is a combination of two linked genes which produce a toxin and an antitoxin. While the toxin is a protein molecule, the antitoxin can be either a protein or an RNA. Normally, both toxin and antitoxin are produced simultaneously and form a complex, with the antitoxin neutralising the toxin, allowing the bacteria to “live happily,” explains Mahavir. But during viral infection, the bacterium uses some factors released by the virus itself to break down this complex, allowing the toxin to poison and kill the bacterial cell, which also ends up killing the virus.
Mahavir’s lab is interested in an unusual TA complex called the type III TA system, where the antitoxin is not another protein, but a stretch of RNA. The genes coding for both toxin and antitoxin are within the same cluster. The toxin gene gets transcribed to RNA and then translated to produce the toxin protein, but the antitoxin gene only gets transcribed to RNA. This antitoxin RNA has a unique appearance – it has a central hairpin-like structure called a pseudoknot, with two straight ends.
Another unusual feature about this TA system is that the toxin protein is actually an enzyme that cuts up its own antitoxin RNA into bits that then bind to the toxin to form the TA complex. Mahavir’s lab has shown that there are at least five types (clusters) of such complexes which can exist in E. coli bacteria. They have also published a detailed crystal structure depicting the tightly-bound toxin-RNA complex in the bacterium.
“In the next set of experiments, we will be working on other unique type III TA complexes in disease-causing bacteria like Staphylococcus and Salmonella,” Mahavir explains. “One of our interests is to understand precisely the role of the pseudoknot in RNA antitoxin in the assembly of this complex.”
Deciphering the structure of such complexes can help target infectious bacteria and pave the way for new antibiotics. “What you have to do,” Mahavir says, “is find a molecule or peptide that can disrupt this TA complex in infectious bacteria, freeing the toxin. And then this toxin will kill the bacteria by itself.”
While research takes up most of his daily routine, Mahavir also makes time for indulging his other passions, like cooking and Taekwondo. “I love my Taekwondo classes,” he elaborates. “I feel that everyone should take up a martial art as a hobby. Its benefits are manifold.”
What keeps him motivated every day is quite simple, he says. “As long as I can devise the next logical experiment, I am happy.”

With input from Malavika P Pillai